Database Discrimination?
Precision medicine often relies on population databases – but this may render it less effective for non-European patients
When is “precision medicine” not precision medicine? When it’s used for patients of non-European descent, a new study from the University of Southern California reveals (1). Ideally, genetic mutations in cancer cells are highlighted in a comparison with normal tissue – but, in many cases, there’s no normal tissue sample available. Genetic information from population databases can serve as a stand-in, but there’s a catch: most of the genomes included in such databases come from individuals of European descent. What does that mean? Variants that are harmless in a given patient may stand out as potentially cancer-causing, simply because the population database lacks the information to identify them as benign.
“A physician could give a treatment that is toxic, ineffective or worse – unnecessarily,” says David Craig, principal investigator and co-director of the Institute of Translational Genomics at USC’s Keck School of Medicine. “This would be the case in the context of clinical decision-making based on tumor sequencing only.” If reported mutations are interpreted as cancer-driving when they are, in fact, inherited and most likely benign, patients might undergo more intensive treatment than necessary, or might not receive the treatment best-suited to their particular disease profile.
“The amount of incorrect inherited information within a precision medicine cancer genomics report is very important, as that speaks towards the precision of the test,” says Craig. “Precision is a part of precision medicine. In research studies attempting to discover new driver mutations and link them to therapy, imprecision lowers the overall chance that a study will yield meaningful new insights.”
The study shows that precision is ancestry-dependent. In some populations, particularly those of European ancestry, the precision is good. In others, it drops precipitously. But the report doesn’t stop there; it goes on to demonstrate analytical approaches to reducing imprecision. How? By deconvoluting normal and tumor tissues from the same sample, taking advantage of the fact that most specimens sent to pathology are not pure tumor to allow comparisons between the two.
So why don’t hospitals routinely collect healthy tissue from cancer patients? Craig says there are many reasons. “Some that are really important, but not frequently discussed, are due to the regulatory uncertainty of explicitly collecting normal specimens, and how it impacts the ability of the physician to act quickly to identify the best therapy.”
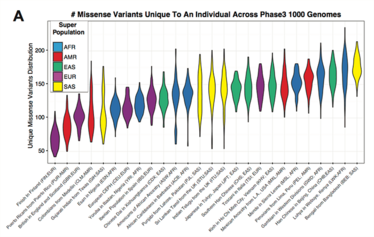
Figure 1. The number of unique missense variants in individual population subtypes. These variants are poorly represented in genomic databases and can lead to false positive results on genomic tests.
In the United States, for example, some state laws require additional genetic counseling prior to conducting tests involving inherited information (2). Regulation around germline testing could add uncertainty to the process and, according to Craig, many view this uncertainty as counterproductive. “Think of it from the perspective of an oncologist working with their patient. The ordering physician may be well aware of different cancer treatments, their effectiveness and how a patient may respond. However, there may be additional laws that require the patient to have genetic counseling before the test is ordered, as there may be incidental findings about family members. For many physicians, introducing regulatory uncertainty about what steps must come before even ordering the test is a major concern.”
But understanding the nature of the problem also suggests a solution. “Our approach identifies ways to separate tumor and normal by recognizing most solid tumors are mixtures. We can use tools to computationally indicate which mutations are from the tumor and which are inherited. We have made those tools available in an open framework (github.com/tgen/LumosVar) that allows the approaches and concepts to be adapted, integrated and validated within future clinical tests.”
LumosVar is not currently a clinical test. It is a research tool and algorithm that its creators hope will lead others to test and validate approaches to deconvolute mixtures. “We hope it enables sequencing of archival samples from diverse populations when requiring a normal means losing diversity,” says Craig. “In our studies, we have seen examples where access to archival tumor specimens is available for African and Asian populations, and we want to make sure that we can maximize the utility of these samples.” It’s vital that our understanding of genetics incorporate as many different populations as possible – because what begins as research on an underserved population may eventually lead to better care for those patients.
- RE Halperin et al., “A method to reduce ancestry related germline false positives in tumor only somatic variant calling”, BMC Med Genomics, 10, 61 (2017). PMID: 29052513.
- National Cancer Institute Cancer Diagnosis Program, “50-state survey of laws regulating the collection, storage, and use of human tissue specimens and associated data for research” (2004). Available at: tp.txp.to/NTRL/library. Accessed November 17, 2017.















