The Inside Story
The Maastricht MultiModal Molecular Imaging Institute (M4I) continues to break new ground in mass spectrometry imaging – providing doctors with better diagnostic tools, and giving us a close-up view of the complex molecular machinery that underpins health and disease.
Breaking boundaries is a key theme of my work, and pivotal to all fields of science. The motivating force of most scientists is to break through boundaries of knowledge – seeing something that no-one has seen before is an awe-inspiring experience. Whether they are building better microscopes to visualize molecules in a cell, or better telescopes to detect far-away stars, scientists have the same drive – they want to see what they cannot yet see.
On a more down-to-earth level, breaking boundaries between disciplines is a crucial facet of our work. We need to make sure that our knowledge crosses the boundaries of our own disciplines, to answer the big questions society faces. Whether it is in life science, food, water or energy, input is needed across the boundary of physics, chemistry, biology and mathematics. All of these different disciplines meet in analytical science, which is why our field lies at the base of so many great discoveries.
I have always been driven by curiosity; I just love figuring out how the world around me works. If the switch on my bike light stops working it’s not enough to simply replace it - I want to understand the problem and try to fix it. The same curiosity that sees me dismantling my bike light also motivates my work, albeit the questions I ask are much bigger! At the Maastricht MultiModal Molecular Imaging Institute (M4I) we seek to visualize fundamental molecular processes, and apply that knowledge to improve human health.
Clinicians often have very sparse information to work with – they are forced to make life-and-death decisions without all the pieces of the puzzle. It’s clear to me that the future of medicine lies in clinicians gaining much more detailed information about the patient, to deliver more personalized treatment, with a better outcome. This personalized medicine – or, as scientific visionary Leroy Hood terms it, personal, predictive, preventive and participatory (P4) medicine – is where I focus my work.
To make personalized medicine a reality, we must find a way to resolve the incredible complexity of the human body and apply it in clinical decision making. And that involves gathering as much information as possible at the genome, proteome and metabolome level and coupling it to disease manifestation, treatment choices and, ultimately, patient outcome. After we gather all of this data, we can start building complex clinical decision-making models. Developments in information technology, machine learning and artificial intelligence have already started to play an increasingly important role in this process, as the sheer volume of the available personal data becomes too daunting to interpret for an individual clinician (or researcher, for that matter). Data scientists will lead clinicians, and will in turn be led by clinicians, analytical scientists and epidemiologists. It’s a nice example of knowledge crossing borders to improve healthcare on many levels.
How do we gather the data needed by modern medicine? For me, mass spectrometry and, more specifically, mass spectrometry imaging (MSI) is central to the endeavor. Mass spectrometry already provides insights into many of the molecular classes found in complex clinical samples, such as blood, urine, cerebrospinal fluid, and many more. Combined with modern chromatographic separation technologies (GC, CE, LC, LC×LC, and so on) mass spectrometry is capable of unraveling the molecular complexity that we need to form the input for our clinical decision-making models. MSI takes this detail to the next level, with analyses performed in the spatial context of cell and tissue.
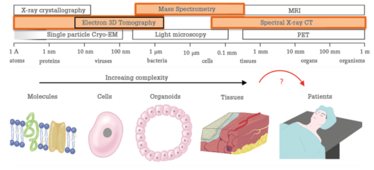
At M4I, we have brought cutting-edge MS-based technologies together with high-end cryo-electron microscopy. In doing so, it becomes possible to image biological processes at multiple scales: a single molecule, the molecule in the context of a cell, the cells in context of diseased and healthy tissue, and that tissue in the context of the patient’s biological system.
Mass Spectrometry Imaging 101
By Shane Ellis, Assistant Professor at M4I.
Mass spectrometry imaging is a molecular imaging technique that exploits a unique feature of every molecule – its weight. By measuring the weight (mass-to-charge ratio) of all the molecules from a small region of a sample we get can determine the spatial locations and concentrations of molecules present in the sample. By sampling many points on a sample we can build detailed images (ion distribution maps) of hundreds of molecules simultaneously. This allows us to see how the presence of certain molecules alters others in the surrounding environment, and how localized chemical processes vary across a complex and heterogeneous sample. MSI involves a variety of techniques, such as pulsed-UV laser irradiation (MALDI), focused ion beam irradiation (SIMS) or charged solvent droplets (DESI). Each method has strengths and weaknesses, and here at M4I we combine all three to help us find answers to complex biomolecular questions.
Mass effort
My group at M4I is pushing the boundaries of spatial resolution in MSI, including developing new tools to resolve molecular structures that have so far proved elusive. We are currently working on combining ion chemistry with MSI to apply imaging in a completely new way (see article) – if successful, this could result in a paradigm shift for the analytical application of mass spectrometry in structural biology.
Throughput is another key area for us. MSI provides orders of magnitude more detailed information for clinical diagnostics than conventional imaging techniques, but the information needs to be available to clinicians quickly. A diagnostic approach that takes hours to complete will not be adopted easily. We are working with the main MS vendors to deliver MSI-based tissue diagnoses to surgeons, pathologists and other healthcare professionals in a matter of minutes (see article), which will facilitate the translation of our work into personalized medicine, ultimately reducing diagnostic and treatment costs.
Whether it’s fundamental research, instrument development or clinical translation, an important bottleneck is our ability to deal with the ongoing data tsunami. We need innovative data sciences and bioinformatics to digest the data as rapidly as we can now generate it (see article).
I believe that the team at M4I has the vision and drive to help move healthcare forward (read more about the work of some of our ‘rising stars’ in the following sections). But to achieve our goals, it’s crucial that our research is embedded in the clinic – for example, our collaboration with nearby academic hospital, MUMC+. M4I has developed several MS-based translational medicine projects together with the MUMC+ Department of General Surgery and Pathology. After three years of building bridges, these projects are now beginning to come to fruition – not only scientifically but also clinically, with the birth of several novel diagnostic assays that are now being validated for patient care. This type of interdisciplinary research is something I am passionate about – it is absolutely crucial to advancing MSI, and science as a whole.
A beautiful example is the recently published (1) result of a collaboration between M4I and Steven Olde-Damink’s group at MUMC+ on non-alcoholic steato-hepatitis (NASH). Using a mass spectrometry-based tissue imaging approach we established a novel classification model for NASH. Importantly, we found that this classification was only possible when the spatial context of the molecules was taken into account; we could not establish the classifier on a tissue homogenate in which the spatial context was lost. This study is a prime example of why MSI is needed for modern medicine.
We are entering a new era of digital pathology, and MSI fits seamlessly with innovative concepts in molecular pathology. In the future, a pathologist will order an MSI assay with similar ease to an immunohistological assay – and examine it on exactly the same platform – allowing diagnosis to be based on much more extensive molecular information. It also offers the combination of targeted and untargeted molecular diagnostics, which will be the true paradigm shift in personalized medicine and molecular pathology.
MSI-based molecular pathology can be combined with MS-based intraoperative diagnostics, as is being done with rapid evaporative ionization mass spectrometry (REIMS), with the “i-Knife” sampling device (see article). Our group has been the first to take all of our molecular imaging information and put it into models that classify tissue during a surgical procedure. We are building the molecular operating room of the future, to improve the quality of care and treatment outcomes of patients.
Scratching the surface
The potential impact of MSI is hard to overstate. Everything we experience, invent, touch, eat and use involves some form of surface chemistry. MSI can help us to better understand all of these surfaces and their chemical interactions with their environment, provided we are careful to ask the right questions and design our experiments in the best way.
The most obvious impact will be in medicine. MSI can be employed to more precisely define a tumor margin on a tissue section, determine the degree of ischemic damage in an organ for transplantation, classify the severity of a disease, and so much more. The pharmaceutical sector will benefit from detailed information on local drug metabolism – researchers will be able to see if a drug reaches a target, without the need for labels that could interfere with its mode of action. Better information on local drug metabolism and pharmacokinetics (DMPK) will be crucial to innovation in drug development.
Food is another area where MSI could make a major impact. For example, we are collaborating with a local organic wine farmer to understand how his method of spraying a microbial extract on his vineyard improves the quality of his plants, his soil, and ultimately his wine. We have scanned leaves from his vineyard throughout the season with DESI-MSI to get a picture of what is happening on the surface of the plant. These type of studies will lead to innovative biological pest control methods and reduce the chemical footprint of the farmer on the environment – only one of the many ways in which MSI will contribute to sustainable agriculture.
The impact of MSI on science and society is already tremendous and can only grow. If the number of published papers is an indication of the impact of MSI, the best is yet to come! Read on to find out more about the young researchers breaking through scientific and technical barriers at M4I.
History of Mass Spectrometry Imaging
There are three main categories of mass spectrometry imaging (MSI): secondary ion MS, ambient MSI and laser-based MSI.
In the sixties, secondary ion mass spectrometry (SIMS) appeared as one of the first surface analysis technologies. Researchers employed energetic ion beams to generate secondary ions that would tell them something about the properties of a surface. At first, they focused on elementary surface composition, but they quickly realized that SIMS could be deployed to study surface chemistry, which piqued the interest of physical chemists. Modern SIMS instruments can study organic surfaces in unprecedented detail. Recent advances have included the implementation of gentle Ar-cluster beams that sputter surface without any organic subsurface damage, allowing us to build full three-dimensional molecular models of single cells. Equally revolutionary was the implementation of tandem mass spectrometry for structural identification, which moved the field from “pretty pictures” of individual m/z values to interpreted biological images. These technologies have found their way into application domains ranging from material sciences, catalysis, forensic sciences, semiconductor sciences, coating technology and, of course, biology and biomedicine.
Ambient MSI was developed when researchers realized that not all samples were suitable for the vacuum of a mass spectrometer. It took until early this century for a suitable ionization technology to be conceived – and desorption electrospray ionization (DESI) is still one of the main ambient imaging technologies for non-vacuum-compatible surfaces. It deploys a supersonic jet of charged droplets that impact the surface and pick up surface molecules. Like SIMS, it involves charged particles that impact a surface, but the desorption and ionization mechanisms are markedly different. The development of DESI resulted in a new field of imaging research and is used to directly study plant surfaces, hydrogels, water-containing polymers, drying paint and bacterial colonies on agar plates, to name just a few. It is also widely used in biomedical tissue imaging, as it requires little to no sample preparation.
Though SIMS and ambient techniques have been valuable, the laser-based technologies have arguably had the biggest impact on MSI. In particular, MALDI-MSI has revolutionized MS-based molecular pathology. The key advantage over the other two technologies is that MALDI-MSI can offer information on a much wider variety of compounds, ranging from metabolites, lipids, peptides, proteins and intact polymer molecules directly from complex surfaces. Even though every molecular class requires a different sample preparation protocol, the breadth of molecular coverage, even within a single class, is still unsurpassed. Spatial resolution has evolved over the years from hundreds of micrometers to just 1–5 micrometers, making the technology compatible with morphological features of interest to pathologists.
All three types of imaging, whether particle- or photon-based, have benefitted from the technological advances in mainstream mass spectrometry. MSI can now be routinely performed on modern hybrid high-resolution instruments such as Fourier transform ion cyclotron resonance mass spectrometry and Orbitrap systems. In addition, developments in time-of-flight (ToF) mass spectrometry have provided new high-throughput approaches that allow us to screen a tissue section in 10–15 minutes, dependent on size and required spatial resolution. These two methods combined – high-throughput MS with high-resolution MS – are the cornerstones of MSI-based clinical diagnostics. At M4I we routinely use them back to back – high-throughput MALDI-ToF-MSI to screen tissues from large patient cohorts, complemented with high-resolution FT-based MSI on selected samples to identify the molecular profiles found. New methods are surfacing fast; three years from now, the MSI field will undoubtedly look very different to today. As more and more disciplines adopt (and adapt) our technologies, I believe we will move from evolutions to revolutions in the years to come.
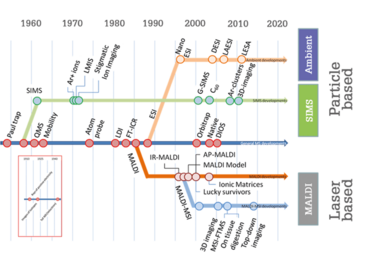
Reproduced from: RMA Heeren, “Getting the picture: The coming of age of imaging MS”, Int J Mass Spectrom, 377, 672-680 (2015).
- K Ščupáková et al., “Spatial systems lipidomics reveals nonalcoholic fatty liver disease heterogeneity”, Anal Chem, 90, 5130–5138 (2018).
- Ron Heeren is the Director of the Maastricht MultiModal Molecular Imaging Institute (M4I) and Division Head of Imaging Mass Spectrometry at Maastricht University, Maastricht, the Netherlands.















