Exploring the Microbiome – with Mice
Rodent models are a useful tool for gaining a better understanding of the human microbiome – but it’s important to understand their limits
At a Glance:
- It’s becoming increasingly clear that the microbiome plays a big role in health and disease – and research in this area is booming
- Many studies are carried out in murine models – but how well do these translate when applied to human health?
- Findings in mice that relate to irritable bowel disease and obesity have translated into humans, but with some important caveats
- As long as researchers are aware of the limitations of mouse models, they will continue to be an important research tool
The role of the microbiome in human health is one of the hottest areas in biomedical research today. A simple PubMed search for “Microbiota and Microbiome” demonstrates a 700 percent increase in the number of relevant publications from 2009 to 2016. Perhaps unsurprisingly, a large number of these studies used murine models. But how well do microbiome findings generated in mouse models translate to human health? It’s crucial that researchers in this area have a clear understanding of the benefits and limitations of their model systems if they wish to extrapolate their findings to humans.
Benefits and caveats
The advantages of using mouse models are clear: homogenous genetics, accessibility to germ-free, mutant and/or transgenic models, a comparable physiology, and a relatively low cost of performing studies. In addition, the use of mouse models has been key in landmark studies determining the role of the gut microbiota in various diseases. But despite the clear benefits, there are caveats affecting translatability that include gross anatomy, compositional differences of the gut microbiota between humans and mice, and environmental factors that may confound studies.
In humans and mice, the majority of the gut microbiota is represented by two key phyla: Bacteroidetes and Firmicutes (1)(2). However, upon closer evaluation, 85 percent of bacterial genera found in the mouse gut microbiota are not present in humans (1). Krych et al. demonstrated that approximately 80 genera are shared between mice and humans (3), with some genera exclusive to each host (4). Despite these differences, profound shifts from control to disease states have been demonstrated and highlight the usefulness of animal models in elucidating the role of the gut microbiota in health.
Two key areas in which mouse models are being used in translational microbiome research are in obesity and inflammatory bowel disease (IBD) – with mixed results.
Obesity studies
Obesity is a serious and growing health concern caused by an imbalance of energy intake and expenditure with contributions from environmental, genetic and societal factors. Starting from the early 2000s, the microbiome has specifically been described as one of the more important risk factors for developing obesity and associated disorders, such as metabolic syndrome and type II diabetes.
The first study linking the process of fat storage to the microbiome was performed in germ-free and microbiota-colonized mice (5). Since then, animal studies exploiting germ-free mice transplanted with microbial communities from obese or lean mice and humans have been instrumental in exploring the role of microbiome in obesity. Key findings from these studies include demonstration of a causal relationship between the microbiome and obesity, altered composition of the “obese microbiome”, and the intriguing observation that the introduction of a “lean microbiome” can reduce obesity and obesity-associated symptoms. Fueled by the evidence in mice, these concepts have been clinically investigated and, in many cases, confirmed in humans (Table 1).
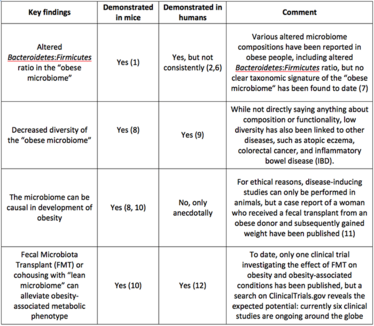
Table 1. Overview of key findings from mouse and human studies on the linkage of the microbiome to obesity and associated disorders.
Mouse models have been a driving force in advancing the obesity-microbiome research field, but translating findings from preclinical studies to something meaningful and actionable in human health remains a challenge. Underpowered human studies have been mentioned as part of the reason, and too much emphasis on small effect sizes, questioning the biological relevance, has been flagged (7). As is so often the case, the myriad of uncontrolled factors in human studies complicates the understanding and interpretation of experimental data. The extensive variation of the microbiome between individuals makes it likely that the “obese microbiome” is an individual concept based on an extremely complex interplay involving at least dietary, genetic, and environmental factors, in addition to microbe-to-microbe synergy and crosstalk. This complexity may explain why a microbial biomarker, or signature, of obesity has not been identified. However, this doesn’t rule out the microbiome as a key contributor.
Better defining IBD
Ulcerative colitis (UC) and Crohn’s disease are the two major types of IBD, and are characterized by chronic relapsing inflammation of the gastrointestinal tract. Underlying both of these conditions is an aberrant immune response to the gut microbiota (13).
The research field has developed multiple mouse models to evaluate specific aspects of IBD, but despite extensive modeling in animals, the precise mechanism through which disease arises has not yet been determined for the majority of patients. As it currently stands, no single mouse model completely recapitulates the pathology of IBD patients. However, many of these developed models remain useful tools when it comes to addressing specific questions relevant to IBD.
More than 790 types of genetic modification in mice have been shown to increase or decrease susceptibility to chemically-induced colitis/ileitis or epithelial barrier dysfunction (14). Additionally, genome-wide association studies (GWAS) have identified more than 160 susceptibility genes in humans, and 20 of these genes have been made into genetically engineered mouse models (GEMs) (14). GEMs are useful in that they can mimic disease phenotypes associated with genetic susceptibility to IBD, but limitations exist in what can be inferred from study results, because these models cannot accurately represent the sum of all genetic differences observed in humans. Furthermore, targeted genes can be involved in numerous pathways, which can cloud the interpretation of results made regarding the associations between gene expression and microbiota.
The substantial differences in the composition of gut microbiota between humans and mice at the genera level make direct translation of experimental results challenging. However, many mouse models of IBD do support a role for the microbiota in experimental colitis (4)(15)(16). Typically, rather than providing translatable linkages between disease and specific microbes, mouse IBD models have demonstrated a role for the gut microbiota as a whole in promoting or preventing disease.
A key finding supporting a role for the resident microbiota in colitis was demonstrated by the observation that following chemical induction of experimental colitis, a significant increase in members of Bacteroidaceae and Clostridium bacteria were detected in the intestines of affected mice (17). Likewise, shifts in the diversity and composition of the gut microbiota of IBD patients are commonly detected (16).
Another finding that demonstrates the role of microbiota in IBD is the resistance of interleukin-10 (IL10)-deficient mice to spontaneous colitis, when they are reared under germ-free conditions (18). This research supports GWAS findings that identify certain IL10 receptor polymorphisms as being associated with the development of early onset IBD (4). In addition, germ-free mice exhibit more severe colitis than conventional mice in a chemically-induced disease model, showing that normal gut microbes may also provide a protective role against experimental colitis (19).
Losing the germs for better modeling
Germ-free mice associated with human microbiota represent a model that has the potential to improve translatability. It’s a powerful tool for researchers because such models can recapitulate a large part of the human gut microbiota composition – 100 percent of phyla, 11 out of 12 classes and ~88 percent of genus-level taxa (4). As mentioned earlier, this model was key in demonstrating the role of the microbiome in obesity. The model was also used in IBD research with the transfer of gut microbiota from UC patients into germ-free BALB/c mice (20). And although none of the humanized mice went on to develop spontaneous colitis, the researchers found that the UC microbiota-associated animals exhibited an increased susceptibility to dextran sodium sulfate (DSS) induced colitis.
However, there are two key caveats to consider when using similar models: i) transplanted microbiota are present in a host that they have not co-evolved with, and ii) the immune system of associated animals has not developed normally from the time of birth with these transplanted microbiota. One partial solution is to vertically transmit microbiota through breeding transplanted mice and performing experiments on their offspring. The diversity and abundance of transplanted gut microbiota is conserved from parental (P) to F1 generation offspring (21). These humanized models have provided valuable insight into the factors that are involved in the progression of both IBD and obesity.
Considerations
Perhaps above all else, mouse models allow for the opportunity to experimentally manipulate and control variables in ways that cannot reasonably be accomplished in human trials. They remain relevant for performing reductionist studies that enable dissection of biological mechanisms and allow for sufficiently powered studies under controlled conditions. Despite the accepted utility of mouse models, and the known anatomical, physiological, and immunological differences between mice and humans, the reductionist approach of mouse studies compared with human studies can also prove to be a limitation: the relationship between the microbiome and the host can appear more complex in humans, and the relevance of single mechanistic pathways revealed through the use of mice may, in some cases, yield results that are not relevant to the human condition. Nevertheless, rodents have already proven their value in our quest to understand the microbiome – providing several solid links between the microbiome and disease that have gone on to be confirmed in humans.
Alexander Maue is portfolio director, Microbiome Products & Services at Taconic Biosciences. Randi Lundberg is a field applications scientist at Taconic Biosciences.
- RE Ley et al., "Obesity alters gut microbial ecology," Proc Natl Acad Sci, 102, 11070–11075 (2005). PMID: 16033867.
- RE Ley et al., "Microbial ecology: human gut microbes associated with obesity", Nature, 444, 1022–1023 (2006). PMID: 17183309.
- L Krych et al.,"Quantitatively different, yet qualitatively alike: a meta-analysis of the mouse core gut microbiome with a view towards the human gut microbiome", PLoS One 8, e62578 (2013). PMID: 23658749
- TL Nguyen et al., "How informative is the mouse for human gut microbiota research?", Dis Model Mech, 8, 1–16 (2015). PMID: 25561744.
- F Bäckhed et al., "The gut microbiota as an environmental factor that regulates fat storage", Proc Natl Acad Sci, 101, 15718–1523 (2004). PMID: 15505215.
- SH Duncan et al., "Human colonic microbiota associated with diet, obesity and weight loss", Int J Obes, 32, 1720–1724 (2008). PMID: 18779823.
- MA Sze, PD Schloss, "Looking for a signal in the noise: revisiting obesity and the microbiome. MBio, 7, e01018-16 (2016). PMID: 27555308.
- PJ Turnbaugh et al., "Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome", Cell Host Microbe, 3, 213–223 (2008). PMID: 18407065.
- E Le Chatelier et al., "Richness of human gut microbiome correlates with metabolic markers", Nature, 500, 541–546 (2013). PMID: 23985870.
- VK Ridaura et al., "Gut microbiota from twins discordant for obesity modulate metabolism in mice", Science, 341, 1241214 (2013). PMID: 24009397.
- N Alang, CR Kelly, "Weight gain after fecal microbiota transplantation", Open Forum Infect Dis, 2, ofv004-ofv004 (2015). PMID: 26034755.
- A Vrieze et al., "Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome", Gastroenterology, 143, 913–916.e7 (2012). PMID: 22728514.
- RJ Xavier, DK Podolsky, "Unravelling the pathogenesis of inflammatory bowel disease", Nature 448, 427–434 (2007). PMID: 17653185.
- A Mizoguchi et al., "Genetically engineered mouse models for studying inflammatory bowel disease", J Pathol, 238, 205–219 (2016). PMID: 26387641.
- J DeVoss, L Diehl, "Murine models of inflammatory bowel disease (IBD): challenges of modeling human disease", Toxicol Pathol, 42, 99–110 (2014). PMID: 24231829.
- KK Gkouskou et al., "The gut microbiota in mouse models of inflammatory bowel disease", Front Cell Infect Microbiol, 4, 28 (2014). PMID: 24616886.
- I Okayasu et al., "A novel method in the induction of reliable experimental acute and chronic ulcerative colitis in mice", Gastroenterology, 98, 694–702 (1990). PMID: 1688816.
- KS Matharu et al., "Toll-like receptor 4-mediated regulation of spontaneous Helicobacter-dependent colitis in IL-10-deficient mice", Gastroenterology, 137, 1380–1390 (2009). PMID: 19596011.
- S Kitajima et al., "Dextran sodium sulfate-induced colitis in germ-free IQI/Jic mice", Exp Anim, 50, 387–395 (2001). PMID: 11769541.
- Z Du et al., "Development of gut inflammation in mice colonized with mucosa-associated bacteria from patients with ulcerative colitis", Gut Pathog, 7, 32 (2015). PMID: 26697117.
- R Lundberg et al., "Microbiota composition of simultaneously colonized mice housed under either a gnotobiotic isolator or individually ventilated cage regime", Sci Rep, 7, 42245 (2017). PMID: 28169374.
Alexander Maue is portfolio director, Microbiome Products & Services at Taconic Biosciences. Randi Lundberg is a field applications scientist at Taconic Biosciences.















