Variant Database Collaborations – for Cancer and Beyond
Is there a better way to sift through the vast quantities of information found in the scientific literature – and still find what’s relevant?
At a Glance
- Keeping genomic data updated with the latest research and clinical findings is valuable to ensure that your patient information is always accurate
- To keep up with the volume of genomic information being produced is too big a task for a single institution, so collaboration is key
- The CIViC, ClinGen and ClinVar databases allow a curated collaborative approach to keeping genomic information up to date
- If widely used, this approach could be integrated with healthcare information to inform clinicians of new indications in old patient data
The already complicated task of organizing and updating scientific literature becomes increasingly more so as information continues to amass at a frightening pace – a challenge that’s especially relevant to cancer genomics. Not only is it important to collate relevant literature for each subject, but also to ensure that any stored information is updated with the latest knowledge. Such a gargantuan task is too much for one group alone to tackle. Here, Obi Griffith and Heidi Rehm discuss how the scientific community at large could offer a solution.
How did you become involved with cancer genomics?
Obi Griffith: I started in bioinformatics. Shortly after college, I worked at Canada’s Michael Smith Genome Sciences Center, which was one of the first big genome centers in Canada – and it was at the forefront of next generation sequencing. We had sequenced genes from humans and many other species, with a strong focus on cancer because of the support from the BC Cancer Agency. Next, I did a post-doc in cancer genomics at Lawrence Berkeley National Laboratory, before moving to the Washington University McDonnell Genome Institute six years ago – again, with a focus on cancer genomics. I work with cancer genome, sequence, and expression data, but I’m also active in bioinformatics, developing software, tools, and databases, so I tend to work on a variety of cancer types.
Heidi Rehm: I’ve been interested in genetics since high school. I majored in molecular genetics and biochemistry at Middlebury College, then went to Harvard for my graduate studies and became more interested in the human disease aspect. I ended up studying genetic hearing loss in my graduate and post-doctoral studies but, soon after, I was hired to start up a new clinical lab, Partners Healthcare Laboratory for Molecular Medicine. Over the course of the last 15 years, it has become apparent that the current model of how we understand and interpret genetic variation is insufficient for the quantity of information being amassed. Clinical research labs cannot maintain the level of deep expertise in all disease areas needed to provide proper, high-quality interpretation for all indications. That’s why I started thinking about a way for us to all work together as a community.
What is the solution?
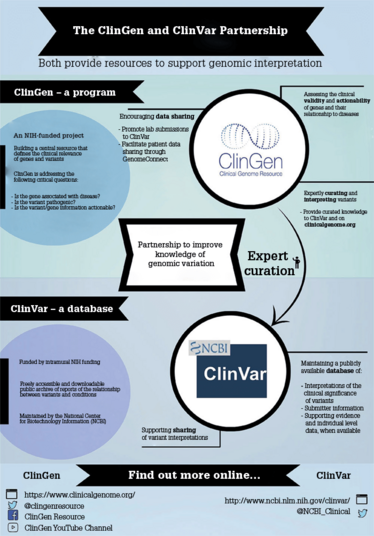
HR: There were people who said, “We need to build a clinical grade database.” At the time, there was the Human Gene Mutation Database, which was based on people going through the literature and finding variants that had been reported in patients. But it had a serious shortfall: there was never really any scrutiny of the evidence base for any of the information. It's a good place to find data in literature reports, but many of the interpretations are incorrect. That was the status quo we were dealing with five years ago, and we weren’t really happy with it. Knowledge constantly changes; even if, at one point in time, all the experts in the world review a variant interpretation as one thing, the next day new information could invalidate it. From that perspective, it almost seems like an impossible scenario to create a curated database able to encompass all the relevant information… Almost! The solution was to let the scientific community become part of the curation process, which is how ClinVar and ClinGen came into being.
ClinVar is a database managed by the National Center for Biotechnology Information, within the National Library of Medicine and National Institutes of Health (NIH). The aim is for people to submit interpretations of variants with supporting evidence, both published and unpublished, to spread the workload of curation across the community and have a means of sharing unpublished data on variants. A star rating system accompanies ClinVar to help users understand the level of review of variant interpretations; ratings go from zero stars (little to no documented methodology) up to four stars (Expert Panel-reviewed).
ClinGen, on the other hand, is a NIH-funded program that aims to develop authoritative resources to define the clinical relevance of genes and variants for use in medicine and research. The ClinGen program has over 570 members – spanning 230 different institutions across the world – participating in working groups. ClinGen forms and approves Expert Panels that review variants submitted to ClinVar from single labs.
OG: Clinical Interpretations of Variants in Cancer (CIViC; civicdb.org) is a newer, NCI-funded database that focuses exclusively on the clinical interpretation of cancer variants. Whereas ClinGen and ClinVar have historically focused more on germline variants, CIViC spotlights somatic variants. Importantly, CIViC is strongly committed to an open data sharing model, with all content in the public domain. Evidence for variant interpretations and assertions is submitted and moderated through a combination of crowd-sourced curation and expert moderation. CIViC is working closely with the ClinGen Somatic Working Group and Global Alliance for Genomic Health (GA4GH) to develop standards for somatic variant assessment.
The pros of using CIViC are that you hope to distribute the work somewhat, and that you achieve more of a community consensus – with transparency. Everything is attributed; you always know who said what and when. Existing, centralized resources are often not accessible to everyone and there is generally no provenance. Moreover, there’s no real mechanism for feedback if you recognize an issue. You could email an author, but it’s not really a direct mechanism for making an improvement to the content.
But like all things, the system isn’t perfect. As it’s distributed, there’s a higher possibility of variable quality, so we editors and moderators need to do a good job of reviewing the content. Additionally, we’re not experts in every subject! So, there’s a possibility for errors. We’re hoping that the transparent model and community allow us to identify such problems quickly.
What are the real-world applications of ClinVar/ClinGen/CIViC?
OG: When you sequence a tumor, you’re sometimes faced with thousands of sequences. Generally speaking, 99 percent of these are unimportant – they may be a symptom of a mutation process, but only a fraction of individual mutations are important. Your real aim is to find the few mutations in these thousands that cause the cancer to grow aggressively – essentially, a needle in a haystack. To make finding the needle easier, the CIViC database gathers knowledge from hundreds of previous patients and published studies that have up-to-date knowledge – and gives us a clue as to which mutations we should be thinking about.
HR: In the cancer setting, tumor versus normal tissue can be an incredibly useful filter to help identify what mutations might be associated with tumor growth and cancer progression, allowing us to exclude most of the variation in the germline and focus in. Unfortunately, as Obi mentioned, there’s almost never just one variant, so we’re still left with the same dilemma: which sequences are passenger mutations that have no role in tumor growth and progression? It’s an ongoing challenge. Sometimes we understand the role of variants, but most of the time, we don’t. That’s where CIViC comes in; it brings together many different somatic cancer databases, doing the same thing we’re doing in the germline space.
OG: CIViC is very much the capstone of the knowledge databases. There are hundreds of thousands of studies on cancer genetic data and sample mutations – and the field is expanding every day. If you’re dealing with the identification of mutations in a particular tumor, you’re going and searching – sometimes in a very manual way – the available literature to help guide your decisions. We understand how laborious that process is, so CIViC was created as a high-level summary of the literature, complete with the potential for clinical colleagues to comment and contribute information based on their experiences – almost like expert-level crowdsourcing.
How is the curation process going to work?
OG: We’re really just summarizing existing knowledge in the literature, so what we’re aiming for is a faithful or accurate representation of evidence presented in the studies. We’re not creating new knowledge; we’re simply synthesizing. When someone submits a new variant interpretation or evidence for certain interpretations, we assess whether it’s a faithful representation of what is being referenced. Then, as not all papers are created equally, we categorize and weight the level of quality of evidence. For example, a report of one anecdotal case is weighted less than a large clinical trial. Much debate and effort go into those judgments of quality, and then we decide how we should synthesize various competing results into one cogent consensus of the current state of belief for that variant.
HR: Within ClinGen, our Expert Panels encourages labs and locus-specific databases to submit all variant interpretations with evidence to ClinVar. The Expert Panel reviews the evidence on each variant and then either approves interpretations or changes interpretation based on expert review. Sometimes that involves aggregating evidence from multiple labs to move interpretations from uncertain significance to classified (pathogenic or benign); other times, it involves resolving differences in variant classifications between labs.
What could the future hold for this kind of database?
HR: A currently unfunded proposal I have is to actually structure the genetic test reports so that variants are clearly mappable to the genome and are able to interface with electronic health records, using the reported ClinVar data as a reference for the variants. As mentioned previously, medical knowledge changes over time. What if a variant that we previously thought was benign becomes pathogenic? If this proposed system were in place, it could update attending physicians and tell them, “This variant has been interpreted by an expert panel with recent information that differs from the original report. You might want to consider re-contacting your patient.” Such high-tier information might be directly used in the healthcare setting in the future.
OG: In building resources like CIViC, ClinVar and ClinGen, we are really trying to tackle how genetics impacts human disease in general. We’re trying to create resources that will eventually allow these approaches to move out of the research setting and into the patient care setting.
Sequencing is becoming quite common. I think the biggest problem coming into the field is the diversity of options. Do you do exome sequencing, whole genome sequencing, or a panel? Do you go with commercial vendors or not? The regular clinician (who’s not a physician-scientist) is unlikely to download a VCF file and interface with databases after they’ve received a sequencing report. Rather, they’re going to depend on the report that’s generated from the clinical diagnostic lab they work with. Right now, there are many disparate variant interpretation resources, but there really isn’t a single go-to source, so you’d probably need a genome atlas, bioinformatician or another kind of expert to help navigate the diverse field of resources at your disposal.
The ClinGen group and others have really championed guidelines where variants get categorized as benign or pathogenic, according to very well-described rules. We’re trying to be inspired by that, but also do things differently. We want to work within the global alliance and other consortia to get the scientific community involved around the world to work together on this problem, because it’s way too big for any one group to solve. Again, that’s why CIViC was built, to help facilitate that collaboration – and we’ve got over 100 contributors to date.
Obi Griffith is Assistant Professor of Medicine and Assistant Director at the McDonnell Genome Institute, St. Louis, USA.
Heidi Rehm is the Chief Genomics Officer in the Department of Medicine at Massachusetts General Hospital and Medical Director of the Broad Institute’s Clinical Research Sequencing Platform, Cambridge, USA.
Obi Griffith is Assistant Professor of Medicine and Assiatant Director at the McDonnell Genome Institute, St. Louis, USA.
Heidi Rehm is the Chief Genomics Officer in the Department of Medicine at Massachusetts General Hospital, and Medical Director of the Broad Institute’s Clinical Research Sequencing Platform, Cambridge, USA.















