
Ghost in the Cell
New software couples computer programming with cell function using the ultimate coding language: DNA
A team of bioengineers at MIT have created a programming language for living cells (1), an advance that could allow even novices to design biological circuits.
The language is called Cello (derived from “cell logic”), and works by converting a text description of a desired computational operation to a DNA sequence. For example, cell circuits can be designed to sense environmental factors, and respond to them by upregulating specified genes.
Every living cell contains a genetic program, encoded in DNA, which controls cell functions via the interactions of a network of regulatory proteins (for example, repressors and activators). To describe how different interactions correlate to function, synthetic biologists use terminology from electrical engineering – for example, one set of interactions might function as a sensor, another as an oscillator.
Synthetic biologists have been using cells as “machines” for some time and have built up a library of genetic parts, such as sensors, biological clocks, and actuators. But the process of designing circuits has traditionally been laborious and required in-depth genetic knowledge. Cello makes the construction of genetic circuits accessible to anyone with basic computer programming skills.
The story behind Cello begins in the late 1980s and early 1990s with lead researcher Christopher Voigt’s teenage hobby of computer programming. “I started off with no interest in biology at all. I wanted to study computer science but my dad suggested a broader degree, so I ended up majoring in chemical engineering,” says Voigt, now a professor of biological engineering at MIT. At college he took no biology, focusing purely on physical science. “What led me to the biotechnology side of things was a bit of serendipity. After picking up some exam results in inorganic chemistry, I was staring at a bunch of comics around a professor’s door. I didn’t notice he was inside, but he spotted me. He called me into his office and started talking about his research in biophysics. By the end of our conversation he had offered me a job. That’s how I got my start.”
From there, Voigt became interested in protein engineering, which led him to synthetic biology. “To me, biology is the ultimate programming language. So when I started my own lab, the team’s focus was on understanding that program,” says Voigt.
Short circuit
Writing Cello was relatively straightforward, says Voigt, but generating functional DNA proved more difficult. The challenge was to get the genetic components – called logic gates by analogy with electronic circuits – to operate in the same way no matter what context the programmer put them in.
At first, using a gate in certain contexts would cause circuit failure, and resolving those kinds of issues took a lot of good engineering, not to mention dogged determination. “Those problems were often because moving the gates in relation to the DNA would alter some of the biochemistry, which would then propagate through the entire circuit. So we had to figure out what was causing these issues, and then create fixes to each one systematically,” says Voigt.
Working with colleagues at Boston University and the National Institute of Standards and Technology, the team carried out a study, published in April 2016, using the system to successfully build biological circuits consisting of up to 12,000 base pairs. An impressive 45 out of 60 circuits designed as part of the study worked the first time they were tested.
It’s a significant achievement, but the team aren’t stopping there; they are continuing work to make the system more robust and flexible. “One big hurdle is the number of gates. Right now we can implement a maximum of 12 – and it gets really difficult after nine – but ideally we’d like 50 or 100 gates in a cell. That would really allow us to unleash the potential of biology,” says Voigt, “The other challenge is moving to different organisms, which really expands what we can do. Each cell has its own idiosyncrasies and nuances to overcome, so we’re trying to create a system that we don’t need to rebuild from scratch for every organism.”
Logic Gates
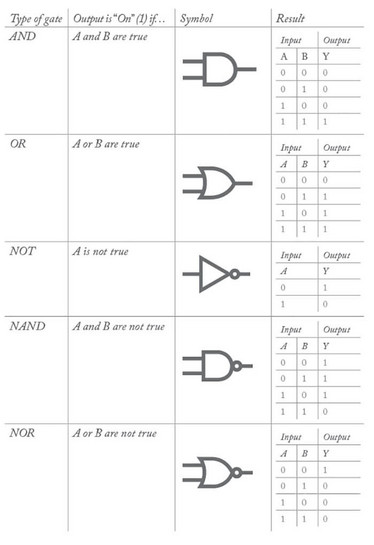
Logic gates are basic building blocks used in digital circuits, which usually have two inputs (A and B) and one output (Y) – all of which are Boolean (they can only be true or false). There are five basic types of logic gates, described below. Cello uses two-input and three-input NOT and NOR gates in the form of repressor proteins.
Ex machina
Cello is based on Verilog, a programming language used for electronic circuits. It allows the user to select from dropdown lists of inputs (sensors) and outputs (actuators), and type in Verilog commands to specify how they should be logically connected. For example, one of the available inputs is the pTET promoter, which switches on or off transcription in response to the presence of the antibiotic tetracycline, while outputs include fluorescent proteins in various colors.
Having compiled your program, just click “run” and Cello automatically calculates the optimum DNA sequence for the circuit. To run the program in E. coli, the researchers synthesized the DNA and inserted it into two plasmids. One plasmid contains the circuit and sensors, while the other encodes the actuator.
“We wanted to implement basic software that would allow users without an understanding of biophysics or genetics to be able to use the logic gates. If a computer is handling that sort of information, it frees the designer up to think about the program they want to run, and not necessarily all the interactions required to implement it,” says Voigt.
The team have made Cello freely available to try online (www.cellocad.org), so biologists and computer scientists alike can try their hand at programming a cell circuit. And the programming code is open source, so anyone can upload new sensors, actuators and “user constraints files”, which define the organism, gate technology, and valid operating conditions. “We wanted to make it freely available so people could be as creative as possible and not worry about the development involved,” says Voigt.
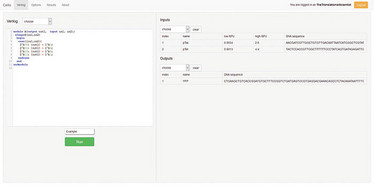
The Cello website – log in, choose inputs and outputs, then connect them using Verilog code (or, like us, use the demo program). Name your creation, and hit “run”.
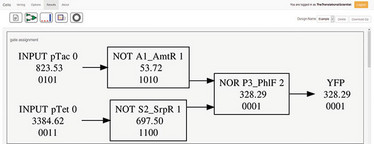
Above is the resulting circuit diagram. In this example, if both tetracycline and lactose are present, the cell will produce yellow fluorescent protein. The results include an optimal genetic layout and plasmid DNA sequence.
Here’s One We Made Earlier
Ctrl Alt Gamete
Synthetic biologists have proposed a range of potential applications for engineered cells, including everything from agriculture to chemical production. Amongst biomedical scientists, the potential for therapeutic applications has sparked interest. Engineered cells could be programmed to navigate to an area of disease and deliver drugs in doses dependent on time, location, or concentration. Another possible application is reprogramming the body’s own microbiome to fight disease.
This research serves as an example of how approaching a field from a different angle, in this case tackling the genome from a computer programming perspective, can lead to advances. “It’s a completely different mindset”, agrees Voigt, “Instead of trying to unpeel biology and all its complexity, the idea is to simplify it all the way down. We pick out the elements that can be implemented by a computer and ignore everything else as much as possible.”
For this reason, Voigt believes that the increasing sophistication of synthetic genomes won’t necessarily teach us a great deal about the workings of genetics in nature. “It’s sort of like how building an airplane doesn’t teach us anything about how birds fly – the principles are totally different. As we’ve gotten better at creating modular systems that can be put together by a computer, we’re getting further and further away from the messy, haphazard genome created by evolution. And in order to make our system designable, we had to implement fixes that look nothing like the genetics you see in biology. Somebody with an expert eye could go into our DNA and instantly recognize that it’s not natural,” says Voigt.
A new breed of bioengineers
With the launch of Cello, Voigt hopes that people with a penchant for computer science might be turned on to the fascinating potential of biological circuits and, like him, discover an interest they never knew they had. “The potential applications are huge and one reason people aren’t getting involved is because it could take years to develop each individual circuit. We thought making it accessible was the best way to speed up the process,” concludes Voigt.
- AAK Nielsen et al., “Genetic circuit design automation”, Science, 352 (2016). PMID: 27034378.
My fascination with science, gaming, and writing led to my studying biology at university, while simultaneously working as an online games journalist. After university, I travelled across Europe, working on a novel and developing a game, before finding my way to Texere. As Associate Editor, I’m evolving my loves of science and writing, while continuing to pursue my passion for gaming and creative writing in a personal capacity.















